Shape-Tunable Plasmonic Gold Nanosensors for Quantitative Circulating Tumor DNA Screening
Abstract:
Cancer has been the leading cause of death worldwide with 1.8 million new cancer cases and over 600,000 deaths in 2020 alone. As the focus of cancer diagnosis has become shifted towards personalized care, molecular diagnostics is a promising method for individualized monitoring of patient tumors, as well as its genetic and phenotypic characteristics. In recent years, liquid biopsy has become a popular method of minimally invasive cancer detection. circulating tumor DNA(ctDNA) is a particularly promising biomarker due to its capability to capture both the genetic and epigenetic changes within a tumor. Moreover, its short, two hour, half-life allows for a representative snapshot of the tumor’s genetic profile. Enrichment and quantitation of ctDNA, however, is a challenge because of its low concentration and the presence of non-tumor specific cell-free DNA in the blood.This study involved designing and fabricating gold nanospheres, nanorods, and nanobipyramids using chemical synthesis to examine the effects of nanoparticle geometry on plasmonic sensitivity. From plasmonic theory, we hypothesized that particles with sharp angular geometries have higher resonance sensitivities due to plasmonic hotspots. We designed the nanosensors through chemical functionalization of the gold nanoparticles, which were conjugated with Peptide Nucleic Acid (PNA) probes for rapid detection of ctDNA. We used a sequentially complementary probe design to capture the G12D variant in Exon 2 of the KRAS gene, a variant extremely relevant to pancreatic and colorectal cancer. We measured the location of the plasmonic resonance peak on the three nanogeometries at five concentrations (0 ng/mL, 25 ng/mL, 50 ng/mL, 75 ng/mL, 100 ng/mL) of ctDNA-spiked healthy patient serum samples.We found limits of detection at 0.6 nanograms/mL for the nanobipyramids and 1.2 nanograms/mL for the nanorods, approaching the clinically relevant range, with the nanobipyramids being two times as sensitive as the nanorods. This was among the first studies comparing plasmonic geometries at nanoscale for quantification of tumor-derived mutations in a clinically relevant scenario. The developed platform may open new opportunities for a wide range of applications from cancer management to drug discovery at molecular scale towards precision healthcare.
Bibliography/Citations:
“Shape Effects of Plasmonic Gold Nanoparticles of Circulating Tumor DNA Screening”, IEEE Sensors, 2020 (co-author)
“Design of Gold Nanorod Based Nanosensors for Circulating Tumor DNA Screening”, 2021 TechConnect World Innovation Conference, June 29-July 1, National Harbor, Maryland, USA (Oral Presentation)
“Rational Design of On-Chip Gold Plasmonic Nanoparticles for Liquid Biopsy”, co-authored manuscript in review
Additional Project Information
Project files
Research Plan:
- Nanoparticle Synthesis and Characterization
Gold nanorods, nanospheres, and nanobipyramids were synthesized in solution. Gold nanospheres were synthesized through stirring and heating chloroauric acid solution in sodium citrate with the molar ratio of 1:10. The gold nanorods and nanobipyramids were fabricated using a seed-mediated method. Throughout both synthesis processes, CTAB was the surfactant to prevent coagulation and serve as a template. The gold seed solution was mixed with a growth solution and left overnight. All nanoparticles were characterized using transmission electron microscopy.
- Development of Electromagnetic Simulation
The resonance features of the synthesized particles were simulated using CST Microwave Studio. The synthesized nanosensors were created in the simulation studio and were simulated as the center of a cube in the chosen refractive index. These spectra were run in both water and in media that represented water glycerol mixtures. The dimensions of the cube were optimized to minimize coupling and boundary effects. We were able to simulate both the shape of the spectrum as well as the sensitivity to changes in bulk refractive index.
- Bulk Refractive Index Testing
To mimic the nanoparticle surface refractive index changes in ctDNA binding, we characterized the particles in water-glycerol mixtures of differing concentrations. After the gold nanoparticles were centrifuged at 6000rpm for 8 minutes, and the supernatant was removed. The collected nanoparticle pellets were incubated in water-glycerol mixtures with varying concentrations(0%, 20%, 50%, 80%, 100%) to examine the response of the surface plasmon peaks to refractive indices (RI). The complete absorbance spectrum was taken from each nanoparticle sample to observe the red shift in the longitudinal peak values due to the changing concentration and refractive index of the mixture. The Tecan Spark 10M Microplate reader from 400nm to 1000nm with a step size of 1nm was used to capture the spectrum.
- Nanoparticle Microfluidic Printing on Chip
The synthesized nanosensors(nanorods, nanospheres, and nanobipyramids) were dispersed on-chip using microfluidic printing. The solution of deionized (DI) water and gold nanoparticles were dispersed into a 96-well plate at a concentration of OD 0.25 (absorbance at peak wavelength). Bare glass slides were functionalized for ten minutes in anhydrous ethanol before thorough drying and microfluidic printing. Then, the nanoparticles were printed using the Carterra Continuous Flow Microspotter onto a glass slide. Optical imaging was used to visualize printed nanoparticle areas.
- Nanoparticle Conjugation and circulating tumor DNA Testing
Gold nanoparticles were washed thoroughly before dispersal onto an activated glass slide.The gold nanoparticles dispersed on chip were conjugated with peptide nucleic acid (PNA) probes complementary to the G12D variant in Exon 2 of the KRAS gene.The nanoparticles were incubated with 2.5mg/mL of DSP (dithiobis succinimidyl propionate) in DMSO for 30 minutes, then coupled to 1mg/mL of the peptide nucleic acid probe(PNABio) in TE buffer for one hour. The sensor was then washed and ready for testing. A FERGIE Integrated Spectrograph (Princeton Instruments) coupled to an optical microscope were used for capturing the spectra. The spectra were then analyzed and processed in MATLAB, which calculated the extinction spectrum and peak location. Peak wavelength location was calculated from the center of mass of peak boundaries.
Questions and Answers
1. What was the major objective of your project and what was your plan to achieve it?
a. Was that goal the result of any specific situation, experience, or problem you encountered?
The objective of my research is to develop novel liquid biopsy technology for early diagnosis and personalized treatment of cancer. I have always been incredibly passionate about science, especially biology and physics towards healthcare applications. I have traveled many places from America, to Asia and to Europe. No matter where I go, I have read stories about cancer mortality rates and their rapid growth in recent years. However, technology development in effective cancer management still lags behind. Personally, I have observed family members diagnosed with the late-stage cancers with limited and non-effective treatment options. These observations resonate with me strongly that early diagnosis tools are much needed to lower mortality rates of cancer. The nanosensor technology developed in this project is such a promising alternative to traditional diagnosis methods, as it is highly sensitive, minimally invasive, and cost efficient.
b. Were you trying to solve a problem, answer a question, or test a hypothesis?
The plasmonic sensors developed in my research serve as a promising tool for circulating tumor DNA (ctDNA) detection towards early cancer diagnosis. Specifically, it tests the hypothesis that the shape and aspect ratio of sensors impact the plasmonic hotspots on biomolecular sensing. Compared to existing nanoparticles and research, this study focuses on analyzing the plasmonic properties of sharp nanogeometric corners, and their effect on ctDNA detection. The synthesized nanosensors were tested on the G12D variant of the KRAS gene, which is implicated in pancreatic cancer. Both experimental data and simulations support my hypothesis that particles with such novel geometries could provide sensitivity enhancement and an interesting system to study LSPR biosensing.
2. What were the major tasks you had to perform in order to complete your project?
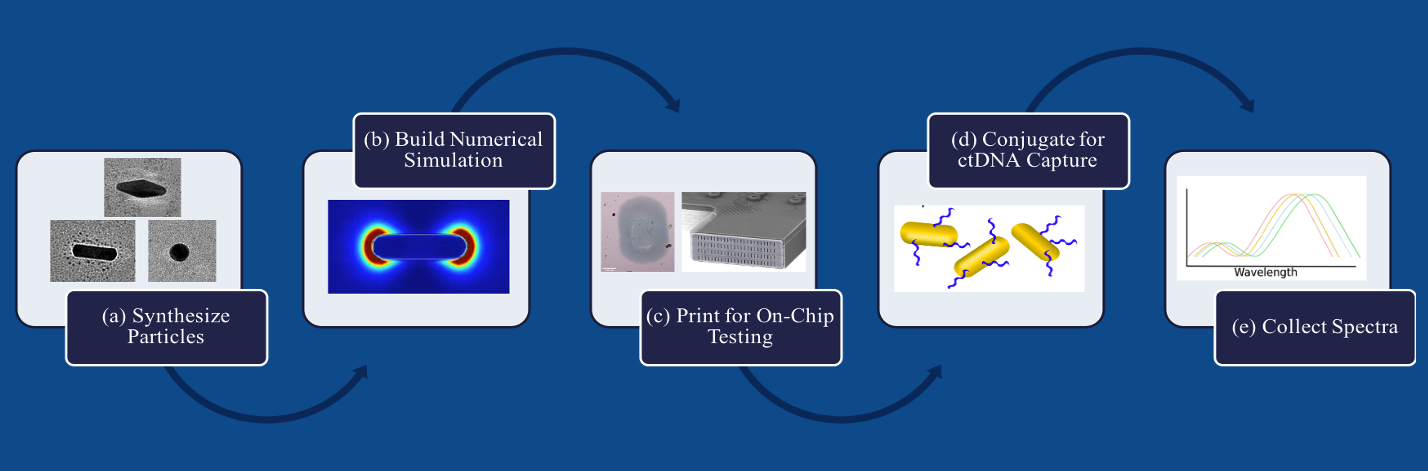
There are three major tasks I accomplished during this research: (a) Design and synthesis of nanoparticles. We synthesized the nanoparticles in the lab and characterized them using the Transmission Electron Microscopy(TEM). (b) PNA probe conjugation. The nanosensor PNA conjugation and ctDNA testing through refractive index variation were completed. (c) Computer simulations of plasmonic performance. As my lab work became virtual under pandemics, I collaborated with other lab members in using a simulation software to analyze the plasmonic sensitivities of the synthesized nanosensors.
3. What is new or novel about your project?
a. Is there some aspect of your project's objective, or how you achieved it that you haven't done before?
This project includes the plasmonic analysis of the nanosensors based on their angular geometries to maximize detection sensitivity. CST simulation software was used for such analysis; as our results proved, simulations were very successful in predicting resonance spectra of the nanoparticles we synthesized in the lab.
b. Is your project's objective, or the way you implemented it, different from anything you have seen?
Common methods of nucleic analysis involve the Polymerase Chain Reaction(PCR) for amplification and sequencing. Due to the low nucleic acid concentration in complex biofluids, PCR process could be lengthy and it requires expensive training. Plasmonic sensors developed in this project offer an “amplification-free”, sensitive, cost efficient, and timely detection device.
c. If you believe your work to be unique in some way, what research have you done to confirm that it is?
This was among the first studies comparing plasmonic geometries at nanoscale for quantification of tumor-derived mutations in a clinically relevant scenario. The significance of the research was recognized by the scientific community through our recent publications. Additionally, my project was recently selected for an oral presentation at the TechConnect International Innovation Conference. The developed platform may open new opportunities for a wide range of applications from cancer management to drug discovery at molecular scale towards precision healthcare.
4. What was the most challenging part of completing your project?
a. What problems did you encounter, and how did you overcome them?
Under the pandemic situation, I had limited access to the lab. However, I was able to continue the study using computer simulations to theoretically analyze the nanosensor plasmonic sensitivity. Such simulations have limitations in predicting arbitrary geometries as shown in real-life scenarios due to, for example, the small inconsistencies that occur in the synthesis process.
b. What did you learn from overcoming these problems?
Modeling and simulations can help provide guidance to sensor design and experiments. To make consistent nanosensor geometries, we can test more parameters throughout the synthesis process. More robust material development and device designs will help verify the theory.
5. If you were going to do this project again, are there any things you would you do differently the next time?
If I were given more lab time prior to the quarantine, I would design and synthesize nanoparticles with more variety of angular geometries. The rich dataset will provide a foundation for future nanosensor development for biomarker screening.
6. Did working on this project give you any ideas for other projects?
As we produce highly sensitive nanosensors through the investigation of plasmonic theory, the on-chip nanobipyramids sets a new platform for portable testing. One idea I have is to print such sensors on a scannable barcode array. This may enable a quick cancer diagnosis test that is readable from a mobile phone scan.
7. How did COVID-19 affect the completion of your project?
During the COVID-19 quarantine, student access to labs was limited. However, I continued to develop my project and collaborate with my mentor to develop new nanogeometries. Specifically, we spent time optimizing the sensitivity through angular nanogeometry. Remotely, we used the CST simulation software to analyze the plasmonic sensitivities and predict plasmonic spectra shape in comparison to the lab data we were able to collect beforehand. There was exemplary agreement in the simulated and experimental data, which shows consistent nanoparticle production and a new method of nanosensor sensitivity prediction using online simulations. Despite the difficult pandemic situation, I keep the research moving forward. We produced a highly-sensitive bipyramid nanosensor, which exhibits twice the sensitivity of the previously synthesized nanorods.