Customized cancer therapy based on the dynamic analysis of the Tumor-Immune-Drug System interaction
Abstract:
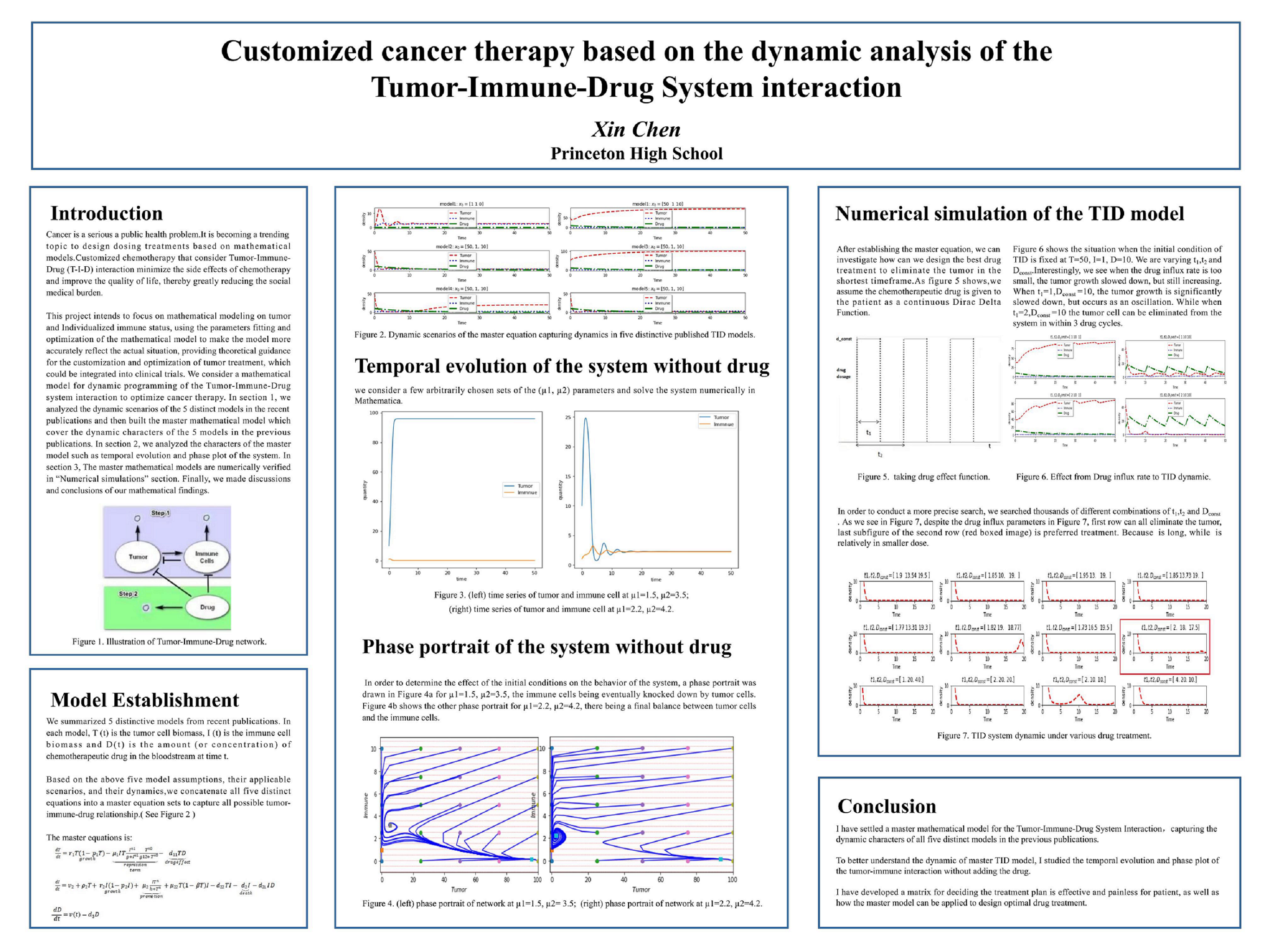
Cancer is a serious public health problem. Despite its subsequent worrying side effects, chemotherapy plays a vital role in cancer therapies. Customized chemotherapy considering Tumor-Immune-Drug (T-I-D) interaction could minimize the side effects and improve the quality of life. Clinically, doctors usually administer chemotherapy at the highest dose that cancer patients can safely tolerate, which often leads to serious side effects and drug resistance. Oncologists believe that therapy adapted to the patient's condition and response may be more effective. The establishment of mathematical models will shed light on the biological evolution of cancer and formulate optimal cancer treatment strategies, guiding clinical trials of adaptive therapy according to the patient's current state. So far, most of the AT policies were developed based on outlier cases that can not be generalized. The problem of optimizing an individual's appropriate schedule and drug dosage remains unresolved.This project intends to focus on mathematical modeling on tumor and individualized immune status, using the parameters fitting and optimization of the mathematical model to make the model reflect the actual situation more accurately, providing theoretical guidance for the customization and optimization of tumor treatment.
I have settled a master mathematical model for dynamic programming of the Tumor-Immune-Drug System Interaction to optimize cancer therapy. I first analyzed all publications about TID model after 2010, and summarized five most representative models that can cover all the models in recent publications. I thoroughly studied the model assumptions and systematic dynamics for all five published models, then built a master model that can cover the dynamic characters of all previously summarized five models. Regardless of the initial model assumption, we can solely use the master equation to describe various TID dynamics and use the master equation to design optimized drug treatment.
To better understand the dynamic of master TID model, I studied the temporal evolution and phase plot of the tumor-immune interaction without adding the drug. Under this model, regardless of the control parameters: µ1, the repression factor from immune cells on tumor; µ2, the promotion factor from tumor on immune cells; the system always settles into a stable solution. But these two parameters will determine the type of steady state. Despite the complexity of the model equations, only two types of behavior are possible – stable spirals and stable nodes. From the dynamic analysis of the model, model are not sensitive to initial conditions, no matter where the tumor or immune cells starts, they will eventually reach a stable state (node/spiral).
Knowing the dynamic of the tumor-immune system is important and instructive for designing future drug treatment. In this project, I have also developed a matrix for determining an effective and less painful treatment plan for patients, as well as how the master model can be applied to design optimal drug treatment.
Keywords: customized cancer therapy / dynamic analysis / Tumor-Immune-Drug System interaction
Bibliography/Citations:
1. Sung, H., et al., Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin 2021;0:1-41
2. Tanay, M.A.L., Medications Used for Cancer, in Understanding Pharmacology in Nursing Practice, P. Hood and E. Khan, Editors. 2020, Springer International Publishing: Cham. p. 393-411.
3. Epstein, R.S., et al., Patient Burden and Real-World Management of Chemotherapy-Induced Myelosuppression: Results from an Online Survey of Patients with Solid Tumors. Adv Ther, 2020. 37(8): p. 3606-3618.
4. Morrison, V.A., Immunosuppression associated with novel chemotherapy agents and monoclonal antibodies. Clin Infect Dis, 2014. 59 Suppl 5: p. S360-4.
5. Chaveli-López, B. and J.V. Bagán-Sebastián, Treatment of oral mucositis due to chemotherapy. J Clin Exp Dent, 2016. 8(2): p. e201-9.
6. Glassman, P.M. and J.P. Balthasar, Physiologically-based modeling of monoclonal antibody pharmacokinetics in drug discovery and development. Drug Metab Pharmacokinet, 2019. 34(1): p. 3-13.
7. Mould, D.R., et al., Developing Exposure/Response Models for Anticancer Drug Treatment: Special Considerations. CPT Pharmacometrics Syst Pharmacol, 2015. 4(1): p. e00016.
8. Abreu, S., et al., Patient-derived ovarian cancer explants: preserved viability and histopathological features in long-term agitation-based cultures. Sci Rep, 2020. 10(1): p. 19462.
9. Urueña, C., et al., Evaluation of chemotherapy and P2Et extract combination in ex-vivo derived tumor mammospheres from breast cancer patients. Sci Rep, 2020. 10(1): p. 19639.
10. Palmer, A.C. and P.K. Sorger, Combination Cancer Therapy Can Confer Benefit via Patient-to-Patient Variability without Drug Additivity or Synergy. Cell, 2017. 171(7): p. 1678-1691.e13.
11. Eriksson, H. and K.E. Smedby, Immune checkpoint inhibitors in cancer treatment and potential effect modification by age. Acta Oncol, 2020. 59(3): p. 247-248.
12. Chan, C.W.H., et al., Novel Strategies on Personalized Medicine for Breast Cancer Treatment: An Update. Int J Mol Sci, 2017. 18(11).
13. Mosa, A.S.M., A.M. Hossain, and I. Yoo, A dynamic prediction engine to prevent chemotherapy-induced nausea and vomiting. Artificial Intelligence in Medicine, 2020. 109: p. 101925.
14. Park, B., et al., Association of white blood cell count with breast cancer burden varies according to menopausal status, body mass index, and hormone receptor status: a case-control study. Sci Rep, 2019. 9(1): p. 5762.
15. Gluzman M, Optimizing adaptive cancer therapy: dynamic programming and evolutionary game theory. Proc. R. Soc. B. 2020,287:20192454. http://dx.doi.org/10.1098/rspb.2019.2454
Additional Project Information
Research Plan:
This project intends to focus on mathematical modeling on tumor and Individualized immune status, using the parameters fitting and optimization of the mathematical model to make the model more accurately reflect the actual situation, providing theoretical guidance for the customization and optimization of tumor treatment, which could be integrated into clinical trials.
Questions and Answers
1. What was the major objective of your project and what was your plan to achieve it?
a. Was that goal the result of any specific situation, experience, or problem you encountered?
b. Were you trying to solve a problem, answer a question, or test a hypothesis?
Five years ago, my loved grandfather was diagnosed with prostate cancer. He became extremely emaciated. I was concerned. He lived with my family since I was little and was an important part of my life. His care involved customized treatment after doctors analyzed his medical records including genetic mutations (such as BRCA2, etc.) They accurately selected molecular targeted drugs (olaparib, PARP inhibitor) for his customized treatment and simultaneously adjusted the chemotherapy regimen with the immune status. My grandfather responded well to the treatment and I was glad to see that healthy grandpa again. Since then, I became interested in research involving customized treatments of cancer. Therefore, I started to learn related knowledge and read relevant articles. This project intends to build a mathematical model for dynamic analysis of Tumor-Immune-Drug System interaction to optimize cancer therapy..
2. What were the major tasks you had to perform in order to complete your project?
a. For teams, describe what each member worked on.
I have done of the work of this project, including literature searching, summarizing representative dynamics scenarios, building up the master mathematical model which cover the dynamic characters of the previous models, analyzing the characters of the master model, numerically verifying the master model.
3. What is new or novel about your project?
a. Is there some aspect of your project's objective, or how you achieved it that you haven't done before?
b. Is your project's objective, or the way you implemented it, different from anything you have seen?
c. If you believe your work to be unique in some way, what research have you done to confirm that it is?
I have built the master equation capturing the dynamic characters of five distinctive published TID systems. This means the master equation is not heavily assumption based. One can imagine, before the master model, to make customized chemotherapy treatment plan, one needs to determine which TID model describes the patient situation the best, then design the treatment using that particular model. Right now, regardless of the initial model assumption, we can solely use the master equation to describe various TID dynamics and use the master equation to design optimized drug treatment..
As it becomes clear with the linearization analysis, the fact that both variables reach a steady state is not simply a special case but rather the only scenario. This result has an important biological relevance because it suggests that neither the tumor, nor the immune cells can grow indefinitely..
I have also developed a matrix for deciding the treatment plan is effective and painless for patient, as well as how the master model can be applied to design optimal drug treatment
4. What was the most challenging part of completing your project?
a. What problems did you encounter, and how did you overcome them?
b. What did you learn from overcoming these problems?
At the beginning, it was difficult for me to understand mathematical models and related biological meanings. I persisted in studying related knowledge and now I can understand mathematical models’ biological meanings. I was not proficient for adjusting the model parameters, so I used manual adjustment. In the future, I will try more systematic and logical debugging methods.
5. If you were going to do this project again, are there any things you would you do differently the next time?
If again, I plan to use the most commonly used techniques for parameter fitting. These include linear least squares, pseudo inverse and eigen analysis, orthogonal least squares, gradient weighted least squares, and bias corrected renormalization.
6. Did working on this project give you any ideas for other projects?
Yes. In this project, I learned that the combination of mathematical models and biological data will turn cancer research into a quantitative and predictable science, linking microlevel information with tumor. I was also exposed to artificial intelligence models. Mathematical model and artificial intelligence are powerful tools of exploration and can provide better solutions to my problems. It became even more intriguing for me after I realized that treating one small aspect may cause a huge change in the whole body. The body not only contains the laws of numerical calculations, but also involves energy exchange and molecular collisions. There are many interesting and unsolved problems in life science that are worth my exploration. How fantastic is this!
7. How did COVID-19 affect the completion of your project?
During the epidemic, my school started online classes, so our teachers assigned a lot more homework than before. In fact, let us get tired of finishing the homework. It is hard to squeeze time to do this research projects that interested me. I am proud to have finally completed it!