Gene Network Analysis of DIO2 and Thyroid Hormone Replacement Therapy
Abstract:
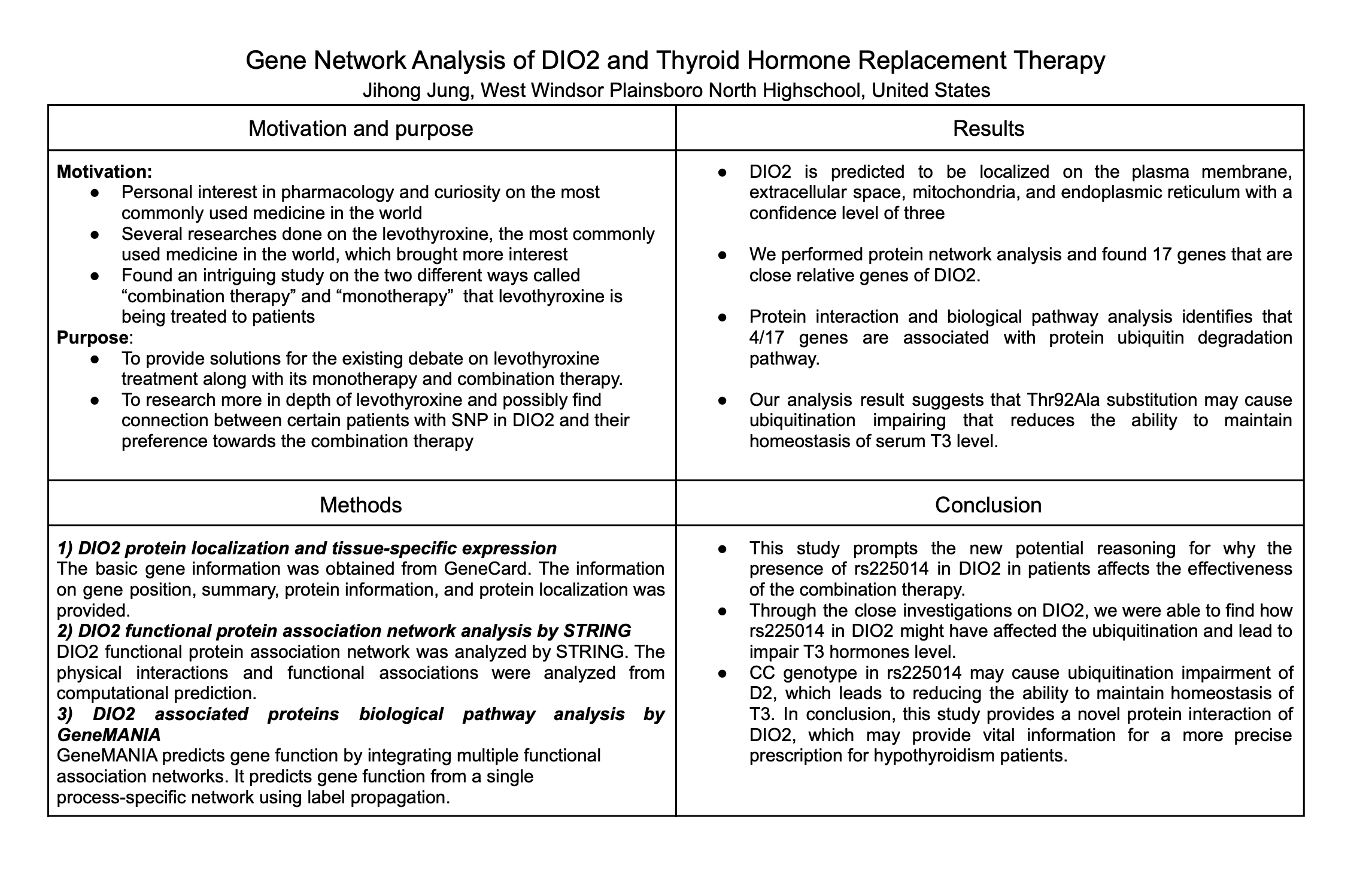
Hypothyroidism is a syndrome in which the metabolic process of the entire body is degraded due to a lack of thyroid hormones. The single nucleotide polymorphism (SNP) information from the DIO2 gene found in the patient's genome is currently used in combination therapy: a treatment for hypothyroidism by providing patients with levothyroxine and liothyronine together. In patients with rs225014 (T→C) transition in DIO2, the conversion from prohormone thyroxine (T4) to bioactive thyroid hormone (T3) proceeds inefficiently even if the level of T4 is increased through Levothyroxine (LT4). However, there is a lack of information to explain how the SNP of DIO2 is linked with the effectiveness of the treatment, and some findings are incomplete and conflicting. In this study, we analyzed the cellular localization of the DIO2 gene, and investigated the tissue-specific gene expression of DIO2 in our body organs. We also conducted a gene network analysis of DIO2 to find the closely related genes to DIO2. Our result shows that the DIO2 gene is closely associated with eight proteins that function as Ubiquitin-dependent proteins. Thus, this study investigated for the first time that DIO2 functions as a homeostasis regulator for thyroid hormones and is closely involved in the proteolytic regulatory system by ubiquitin. The findings provide more information on DIO2 gene function, which can be applied to the development of personalized treatments for hypothyroidism, suggesting that the drugs can be prescribed more precisely and accurately.Bibliography/Citations:
[1] 1. Chiovato, L., Magri, F., and Carlé, A. (2019). Hypothyroidism in context: where we’ve been and where we’re going. Adv. Ther. 36, 47–58.
[2] Hughes, M., Inglese, J., Kurtz, A., Andalibi, A., Patton, L., Austin, C., Baltezor, M., Beckloff, M., Sittampalam, S., Weingarten, M., et al. (2004). Early Drug Discovery and Development Guidelines: For Academic Researchers, Collaborators, and Start-up Companies. In Assay Guidance Manual, G. S. Sittampalam, N. P. Coussens, H. Nelson, M. Arkin, D. Auld, C. Austin, B. Bejcek, M. Glicksman, J. Inglese, P. W. Iversen, et al., eds. (Bethesda (MD): Eli Lilly & Company and the National Center for Advancing Translational Sciences).
[3] Chen, Y., and Tai, H.-Y. (2020). Levothyroxine in the treatment of overt or subclinical hypothyroidism: a systematic review and meta-analysis. Endocr. J. 67, 719–732.
[4] Wiersinga, W.M. (2019). T4 + T3 combination therapy: any progress? Endocrine 66, 70–78.
[5] Vignal, A., Milan, D., SanCristobal, M., and Eggen, A. (2002). A review on SNP and other types of molecular markers and their use in animal genetics. Genet. Sel. Evol. 34, 275–305.
[6] Park, E., Jung, J., Araki, O., Tsunekawa, K., Park, S.Y., Kim, J., Murakami, M., Jeong, S.-Y., and Lee, S. (2018). Concurrent TSHR mutations and DIO2 T92A polymorphism result in abnormal thyroid hormone metabolism. Sci. Rep. 8, 10090.
[7] Panicker, V., Saravanan, P., Vaidya, B., Evans, J., Hattersley, A.T., Frayling, T.M., and Dayan, C.M. (2009). Common variation in the DIO2 gene predicts baseline psychological well-being and response to combination thyroxine plus triiodothyronine therapy in hypothyroid patients. J. Clin. Endocrinol. Metab. 94, 1623–1629.
[8] Wouters, H.J.C.M., van Loon, H.C.M., van der Klauw, M.M., Elderson, M.F., Slagter, S.N., Kobold, A.M., Kema, I.P., Links, T.P., van Vliet-Ostaptchouk, J.V., and Wolffenbuttel, B.H.R. (2017). No Effect of the Thr92Ala Polymorphism of Deiodinase-2 on Thyroid Hormone Parameters, Health-Related Quality of Life, and Cognitive Functioning in a Large Population-Based Cohort Study. Thyroid 27, 147–155.
[9] Safran, M., Dalah, I., Alexander, J., Rosen, N., Iny Stein, T., Shmoish, M., Nativ, N., Bahir, I., Doniger, T., Krug, H., et al. (2010). GeneCards Version 3: the human gene integrator. Database (Oxford) 2010, baq020.
[10] Szklarczyk, D., Gable, A.L., Lyon, D., Junge, A., Wyder, S., Huerta-Cepas, J., Simonovic, M., Doncheva, N.T., Morris, J.H., Bork, P., et al. (2019). STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613.
[11] Warde-Farley, D., Donaldson, S.L., Comes, O., Zuberi, K., Badrawi, R., Chao, P., Franz, M., Grouios, C., Kazi, F., Lopes, C.T., et al. (2010). The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 38, W214-20.
[12] Binder, J.X., Pletscher-Frankild, S., Tsafou, K., Stolte, C., O’Donoghue, S.I., Schneider, R., and Jensen, L.J. (2014). COMPARTMENTS: unification and visualization of protein subcellular localization evidence. Database (Oxford) 2014, bau012.
[13] Pontén, F., Jirström, K., and Uhlen, M. (2008). The Human Protein Atlas--a tool for pathology. J. Pathol. 216, 387–393.
[14] Silva, J.F., Ocarino, N.M., and Serakides, R. (2018). Thyroid hormones and female reproduction. Biol. Reprod. 99, 907–921.
[15] Gereben, B., Goncalves, C., Harney, J.W., Larsen, P.R., and Bianco, A.C. (2000). Selective proteolysis of human type 2 deiodinase: a novel ubiquitin-proteasomal mediated mechanism for regulation of hormone activation. Mol. Endocrinol. 14, 1697–1708.
[16] Dentice, M., Bandyopadhyay, A., Gereben, B., Callebaut, I., Christoffolete, M.A., Kim, B.W., Nissim, S., Mornon, J.-P., Zavacki, A.M., Zeöld, A., et al. (2005). The Hedgehog-inducible ubiquitin ligase subunit WSB-1 modulates thyroid hormone activation and PTHrP secretion in the developing growth plate. Nat. Cell Biol. 7, 698–705.
[17] Meulenbelt, I., Min, J.L., Bos, S., Riyazi, N., Houwing-Duistermaat, J.J., van der Wijk, H.-J., Kroon, H.M., Nakajima, M., Ikegawa, S., Uitterlinden, A.G., et al. (2008). Identification of DIO2 as a new susceptibility locus for symptomatic osteoarthritis. Hum. Mol. Genet. 17, 1867–1875.
Additional Project Information
Research Plan:
1) DIO2 gene general information will be collected
The basic gene information will be obtained from GeneCard. The information on gene position, summary, protein information, and protein localization was provided. It provides a database of human genes that provides concise genomic-related information with all known and predicted human genes. It also provides genomic, proteomic, transcriptomic, genetic, and functional information on all known and predicted human genes.
2) Protein network analysis will be performed using a STRING database.
It is a database of known and predicted protein-protein interactions. The physical interactions and functional associations were analyzed from computational prediction. The interaction database was constructed based on the five main sources: genomic context prediction, high-throughput lab experiments, co-expression, automated text mining, previous knowledge in the database.
3) Analyze the biological pathway of DIO2 using GeneMANIA
GeneMANIA predicts gene function by integrating multiple functional association networks. It predicts gene function from a single process-specific network using label propagation. It provides genome-wide predictions that achieve an accurate seed gene list without relying on a pre-specified association network.
Questions and Answers
1. What was the major objective of your project and what was your plan to achieve it?
The major objective of my project was to find a possible solution to a current debate on the effectiveness of combination therapy of levothyroxine to certain patients. In order to achieve this goal, in-depth research was needed in order to find connections between hypothyroidism patients and the combination therapy. Since the topic has been issued several times, there were many sources that I was able to use in order to develop my understanding and to find new possibilities.
a. Was that goal the result of any specific situation, experience, or problem you encountered?
Hypothyroidism is a thyroid disease that does not produce sufficient levels of thyroid hormone. One of my family members suffers from this disease. Therefore, I was interested in the medication provided for hypothyroidism patients and found an interesting link between DIO2 gene and the prescription preference used in the hospital.
b. Were you trying to solve a problem, answer a question, or test a hypothesis?
I was trying to solve a problem
2. What were the major tasks you had to perform in order to complete your project?
I have done in-depth research in order to gather sufficient knowledge as well as to find new potentials.
a. For teams, describe what each member worked on.
N/A
3. What is new or novel about your project?
My study suggests a new potential ability of DIO2 is not only as a transference between the T4 hormones and T3 hormones but also as the proteolytic regulator with ubiquitin.
a. Is there some aspect of your project's objective, or how you achieved it that you haven't done before?
By focusing on the closely related genes with DIO2, I found out they all have the involvement of ubiquitin. This finding provided a new potential ability of DIO2, thus a new pathway that future researchers can utilize to find the answer to my research question.
b. Is your project's objective, or the way you implemented it, different from anything you have seen?
My objective focuses on the genetic interaction and protein interaction of DIO2. This objective was unique because gene network analysis has not been studied before.
c. If you believe your work to be unique in some way, what research have you done to confirm that it is?
A close relationship between DIO2 and relative genes was found in this research. Since these genes were closely related to ubiquitin-dependent protein degradation, this provides new insight into how DIO2 is involved in regulating thyroid hormone levels in the human bloodstream. 4. What was the most challenging part of completing your project?
a. What problems did you encounter, and how did you overcome them?
Not much information was available for finding a link between DIO2 SNP and ubiquitin-protein degradation activity. I overcome this problem by searching for many research papers.
b. What did you learn from overcoming these problems?
Even though I have been faced with limited information to move into the next step of my research, finding a piece of detailed information on the previously published research papers provides valuable information.
5. If you were going to do this project again, are there any things you would you do differently the next time?
I would like to do more research on the relationship between DIO2 and ubiquitin in order to provide the answer to my research question which I could not do this time.
6. Did working on this project give you any ideas for other projects?
I would like to do more research projects on more medicines that have such varying treatments as levothyroxine.
7. How did COVID-19 affect the completion of your project?
The accessibility to wet experiments was limited due to COVID-19.